Oct 3, 2008
Proteopedia
Sep 24, 2008
Web link reference manager
Useful one check it out this http://www.connotea.org/ and sign up (similar to mail sign up) and make use of it.
Sep 23, 2008
Vadlo - Search engine for research scholar
Sep 12, 2008
Protein crystallography
Check it out this site, if you are interested in protein crystallography http://proteincrystallography.org/
Aug 30, 2008
AIGore Nobel Lecture
http://nobelprize.org/nobel_prizes/peace/laureates/2007/gore-lecture_en.html
Aug 26, 2008
The softer side of science
Mastering soft skills helps master one's career.
Scientists should shield themselves from discouraging events and develop a 'frustration tolerance' for paper and grant rejections, criticism by well-meaning colleagues and the depressing tedium of data collection. Then there's hypermotivation. To avoid burn-out, try relaxation (sports, yoga, meditation) and a healthy social life. Just as learning requires a quiet consolidation phase to store material in long-term memory, success in science needs intermittent silence.
Personality traits cannot usually be changed, but there are ways to improve one's disposition in the lab. Perfection, for example, can only be expected in pure mathematics or fairy tales. Beware of the 80–20 rule: for perfection, the last 20% of a task may take 80% of the effort — not a wise choice if you want to be productive. Worse, perfectionism is a sure path to leaving projects or papers incomplete.
A related trap is failure to bring a project to publication. A finished experiment may satisfy your curiosity, but data are only of value to your CV and the rest of the world if published. The drive to completion is healthy, if you wish to succeed in science. Many of us have papers that are 90% finished but never submitted. Consider the time you have already invested, how little is left to do and how much effort it would take to get to the same point with another project.
Writer's block can be a major challenge: sitting in front of a blank page, lacking the wherewithal to start writing. This may be linked to the perfectionist trap ("I do not write unless my text is perfect"), but it could also be the result of a lack of ideas, of writing ability or of self-confidence. It is usually the first 10 minutes of writing that are hardest. Try to start writing without worrying about the presentation or structure.
Scientific leadership qualities — exhibiting responsibility, flexibility and trust — are also essential in all aspects of science.
These seemingly simple tasks can be taxing. But improving soft skills is a critical element of science success.
From Nature magazine http://www.nature.com/naturejobs/2008/080529/full/nj7195-694b.html#top
Jul 24, 2008
retracted paper from india
I have gone through this paper recently which is retracted from India
RETRACTED: A computational docking study for prediction of binding mode of diospyrin and derivatives: Inhibitors of human and leishmanial DNA topoisomerase-I
Bioorganic & Medicinal Chemistry Letters, Volume 17, Issue 18, 15 September 2007, Page 5281
Sandeep Chhabra, Pooja Sharma, Nanda Ghoshal.
Reference : Bioorganic & Medicinal Chemistry Letters
Jun 7, 2008
Gromacs-tutorial
http://md.chem.rug.nl/education/mdcourse/MDpract.html
May 22, 2008
Research article with reviewers comment in Biologydirect
Various hypotheses on MHC evolution suggested by the concerted evolution of CD94L and MHC class Ia molecules. Biology Direct 2006, 1:3doi:10.1186/1745-6150-1-3
Profiles in science
This is the biographies of some scientist. It is interesting. Have a look at it.
How to setup a default application for a file(like word with Microsoft word)
text files with gedit or nedit
pdf files with acroread or xpdf
xml files with gedit or nedit
For example
I want all "xml" file types to openwith gedit (text editor).
Right click the file and select "Open with" "other application" you will get window like this (shown below). If you want to choose different editor choose the file types available.

Select the file type you want. I have selected gedit (text editor)

Now i can open all the "xml"(any application) files with gedit text editor
May 19, 2008
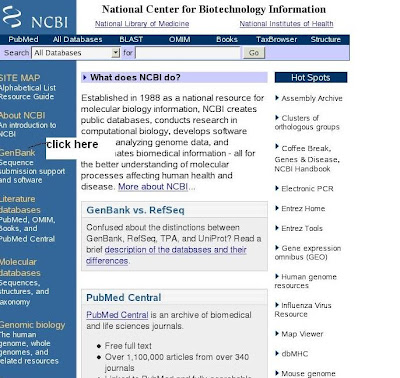
How to submit sequence in genbank (NCBI)
 2)
2)
3)
 4)
4)
5)
1) Open the NCBI home page (click genbank submission hyperlink)
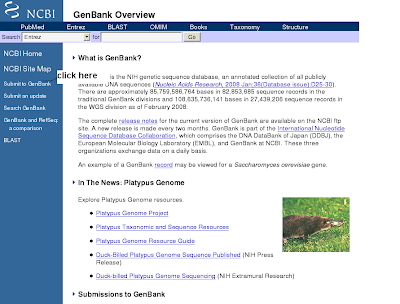
2) You will get genbank overview(click submit to genbank)
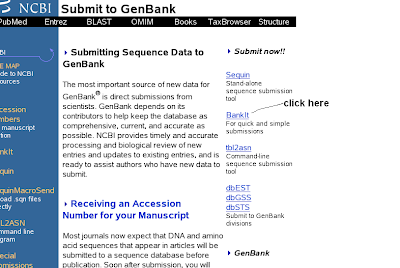
3) You will get submit to genbank (clikc the right side BankIt)
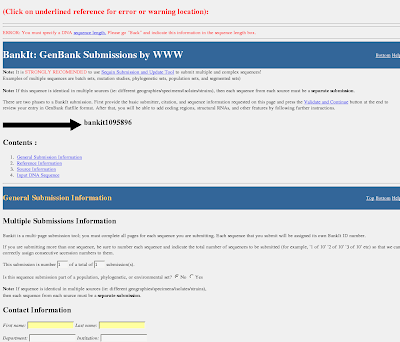
4) You will get BankIt submission form (enter sequence length)
5)You can fill the BankIt sequence submission form
May 15, 2008
INSTALl-postgresql in linux
This is a little more complicated than installing Apache.
The best bet is to install from packages since this will make things easier, both now and for upgrades.
If you decide to install from source code, you'll need to create a user and compile the source.
Create a PostgreSQL user (you'll need root access). This is who owns the system. I use the "postgres" user and group, you can use something different but it's best to keep it related to the package.
See your system documentation on how to do this (hint - man useradd). Give the user a home directory of /usr/local/pgsql as this is where PostgreSQL defaults to when it installs.
Once all that is done, untar the source:
$ tar -zxf postgresql-
or
$ tar -xjf postgresql-
Jump into the directory:
$ cd postgresql-
(replace
Again, configure has a lot of help:
$ ./configure --help
We'll just install into /usr/local/pgsql and leave the rest as-is:
$ ./configure --prefix=/usr/local/pgsql
Next we have to make it (this will take a while, it takes longer than Apache):
$ make
You will get a line:
All of PostgreSQL successfully made. Ready to install.
when everything is finished.
You'll need to log in as root to do the rest:
$ su -
Go back to the postgresql directory and do:
# make install
PostgreSQL doesn't allow root to start up the server for security reasons, so next you'll have to change the owner of the files:
# chown -R postgres:postgres /usr/local/pgsql
Change to the postgres user:
# su - postgres
Then, change to /usr/local/pgsql:
$ cd /usr/local/pgsql
We need to initialize a database cluster:
$ bin/initdb -D ./data
This will create a /usr/local/pgsql/data directory and initialize it ready for startup and use.
As usual, you can view options by using the --help switch:
$ bin/initdb --help
Then you can start:
$ bin/pg_ctl -D ./data -l data/logfile start
Now that postgresql is started, we need to create a database and another user before we go any further.
You should use a separate database for each of our projects, it's not essential though - it just makes things a little cleaner and easier to understand.
You should also use separate users for each database. This keeps everything separate and 'project a' won't be able to modify any of 'project b's data.
To create a new user in postgresql, it's pretty simple:
$ /usr/local/pgsql/bin/createuser
and follow the prompts - pretty simple :)
The new user should not be able to create new databases or create new users.
To create a database, it's a little different.
$ /usr/local/pgsql/bin/createdb --owner=username databasename
We have to set the owner of the database when we create it, otherwise the 'postgres' user owns it and then we have to grant access to allow our new user to access it.
Now that that's all done, log out of the postgres user, so we're back as root. In the postgres directory, the contrib/start-scripts/ directory has a few system start up scripts
Useful installation instruction from http://www.designmagick.com/article/2/Starting-Out/Installing-PostgreSQL
This is a little more complicated than installing Apache.
The best bet is to install from packages since this will make things easier, both now and for upgrades.
If you decide to install from source code, you'll need to create a user and compile the source.
Create a PostgreSQL user (you'll need root access). This is who owns the system. I use the "postgres" user and group, you can use something different but it's best to keep it related to the package.
See your system documentation on how to do this (hint - man useradd). Give the user a home directory of /usr/local/pgsql as this is where PostgreSQL defaults to when it installs.
Once all that is done, untar the source:
$ tar -zxf postgresql-
or
$ tar -xjf postgresql-
Jump into the directory:
$ cd postgresql-
(replace
Again, configure has a lot of help:
$ ./configure --help
We'll just install into /usr/local/pgsql and leave the rest as-is:
$ ./configure --prefix=/usr/local/pgsql
Next we have to make it (this will take a while, it takes longer than Apache):
$ make
You will get a line:
All of PostgreSQL successfully made. Ready to install.
when everything is finished.
You'll need to log in as root to do the rest:
$ su -
Go back to the postgresql directory and do:
# make install
PostgreSQL doesn't allow root to start up the server for security reasons, so next you'll have to change the owner of the files:
# chown -R postgres:postgres /usr/local/pgsql
Change to the postgres user:
# su - postgres
Then, change to /usr/local/pgsql:
$ cd /usr/local/pgsql
We need to initialize a database cluster:
$ bin/initdb -D ./data
This will create a /usr/local/pgsql/data directory and initialize it ready for startup and use.
As usual, you can view options by using the --help switch:
$ bin/initdb --help
Then you can start:
$ bin/pg_ctl -D ./data -l data/logfile start
Now that postgresql is started, we need to create a database and another user before we go any further.
You should use a separate database for each of our projects, it's not essential though - it just makes things a little cleaner and easier to understand.
You should also use separate users for each database. This keeps everything separate and 'project a' won't be able to modify any of 'project b's data.
To create a new user in postgresql, it's pretty simple:
$ /usr/local/pgsql/bin/createuser
and follow the prompts - pretty simple :)
The new user should not be able to create new databases or create new users.
To create a database, it's a little different.
$ /usr/local/pgsql/bin/createdb --owner=username databasename
We have to set the owner of the database when we create it, otherwise the 'postgres' user owns it and then we have to grant access to allow our new user to access it.
Now that that's all done, log out of the postgres user, so we're back as root. In the postgres directory, the contrib/start-scripts/ directory has a few system start up scripts
Useful installation instruction from http://www.designmagick.com/article/2/Starting-Out/Installing-PostgreSQL
May 13, 2008
NCBI Resource Locator
The NCBI Resource Locator provides stable, uniform addressing for NCBI content, making it easy to link to individual records. Some NCBI resources also provide services (like search) through these URLs.
How does it work?
Each URL has the form
http://view.ncbi.nlm.nih.gov///
Where:
* is an NCBI resource (e.g., pubmed, gene, nucleotide, etc.)
* is the action to perform (e.g., search, get, etc.). If is missing, the default verb "get" is used.
* is data used by the action to perform the request
Some examples:
* http://view.ncbi.nlm.nih.gov/pubmed/12345
Show the PubMed record with pmid 12345
* http://view.ncbi.nlm.nih.gov/pubmed/search/cancer
Search PubMed for "cancer"
* http://view.ncbi.nlm.nih.gov/gene/search/human+p53
Search PubMed for "human p53"
* http://view.ncbi.nlm.nih.gov/homologene/search/dystrophin
Search Homologene for "dystrophin"
Reference http://view.ncbi.nlm.nih.gov/
unzip file(bz2,tar) files
use the command "tar -cvf filename.tar" for unzip the tar files
use the command "tar -xzvf filename.tar.gz for unzip the tar.gz files
How to transfer files using SCP in linux
scp filename username@hostname(or ip address):/home/user/ (path)
for eg, in my homology machine i use
scp filename suresh@homology:/home/suresh/